News Release
UC San Diego Computer Scientist Plays Major Role in $25M Cancer Grand Challenges Project
Vineet Bafna will join a team of international researchers investigating extrachromosomal DNA, a major driver of tumor evolution
 |
| Vineet Bafna, a professor in the UC San Diego Department of Computer Science and Engineering, is a key player on the $25M Cancer Grand Challenge Project. |
June 16, 2022-- University of California San Diego computer scientist Vineet Bafna is part of a team of world-class researchers that has been awarded a five-year, $25 million Cancer Grand Challenges grant to learn how the destructive genetic lesion extrachromosomal DNA (ecDNA) influences numerous cancers and to identify possible therapies.
Cancer Grand Challenges is a global research funding program created by the United States National Cancer Institute (part of the National Institutes of Health) and Cancer Research UK. The group funds multidisciplinary research teams to solve some of medical science’s thorniest challenges – in this case, ecDNA, a major driver of tumor evolution. As part of a $500 million initiative, Cancer Grand Challenges announced four teams, including the ecDNA group (called eDyNAmiC), on June 16.
The team will investigate the mechanisms that drive ecDNA—small, circular pieces of genetic information that allow cancer cells to rapidly evolve and become resistant to even the most potent anti-cancer treatments. These genetic anomalies appear in about a third of cancers, promote aggressive tumor behavior and generate poor patient outcomes.
“In many cancers, DNA replication and repair machinery is so dysregulated that we begin to see gross abnormalities in the genome; it is no longer a stable molecule,” said Bafna, a professor in UC San Diego’s Department of Computer Science and Engineering and a faculty member in the university’s HalıcıoÄŸlu Data Science Institute. “Fragments of genetic material break away from their native locations on chromosomes, forming small, circular molecules that are capable of independent regulation.
“These breakaway fragments can duplicate tumor-promoting oncogenes, making 20, 50, maybe 100 copies, helping cancers become more aggressive and resist treatments. While their existence has been known for nearly 80 years, their prevalence in cancer was not fully appreciated until recently,” Bafna said.
The research team
Bafna will be part of a research all-star team led by Paul Mischel, MD, professor of pathology, Stanford Medicine. The group combines expertise in cancer genomics, yeast genetics, epigenomics, patient tracking, machine learning and mathematical modeling, immunobiology, chemistry and other disciplines to interrogate ecDNA’s role in cancer and develop new therapeutic options against it.
“We’re poised to transform the collective understanding of many aggressive forms of cancer and provide new insight into how to diagnose, monitor and treat patients for whom current therapies do not work,” said Mischel.
For this project, Bafna will help spearhead the computational genomics analyses. He has been working for years with Mischel and others to develop mathematical and computational techniques to identify ecDNA and other structural changes in chromosomes.
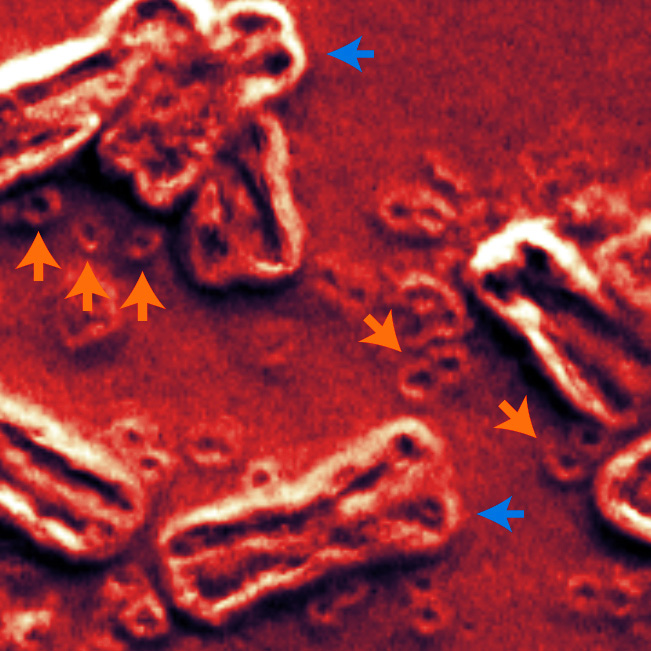
For example, his group maps short (150 base pair) ecDNA fragments to a reference genome to look for duplications, as well as evidence of rearrangements linked to tumor genomes. Using optical mapping technology, they seek to clarify the complex interrelationships between these rogue DNA elements and remaining chromosomes. In a complementary approach, Bafn’s lab has pioneered deep learning methods to detect and quantify ecDNA in cellular images.
“If a region is sampled about 30 times in a normal genome, and 300 times in the cancer genome, you know it has been amplified by a factor of ten,” said Bafna. “However, understanding the architecture of the rearranged genome is more fundamental; it explains how the regulatory circuit is rewired in ecDNA, and if new enhancer elements have been hijacked to further dysregulate oncogene expression.”
This work is becoming increasingly important because ecDNAs are, unfortunately, quite common. Bafna and colleagues have found them in lung and kidney cancers, melanomas and glioblastomas. Duplications were particularly high in glioblastomas, the deadliest type of brain cancer. So far, they’ve found ecDNA in 40% of the tumors they’ve investigated.
“These structural variations are both incredibly important and understudied,” said Bafna. “Now, we can begin to fill in the knowledge gap on ecDNA origin, function and evolution and find ways to counteract it.”
The eDyNAmiC team brings together researchers from 11 institutions across the US, UK and Germany: Stanford University and Sarafan ChEM-H; University of California San Diego; New York University Langone Health; University of Texas Southwestern Medical Center; Scripps Research; Queen Mary University London; University College London; Max Delbrück Center for Molecular Medicine and Charité Berlin; Fred Hutchinson Cancer Research Center; University of Cambridge; Jackson Laboratory for Genomic Medicine.
Media Contacts
Katie Ismael
Qualcomm Institute
858-246-5898
kismael@eng.ucsd.edu
Joshua Baxt
Computer Science and Engineering Writer
000-000-0000
none@none.none